Create a generalized Venn Diagram analog for sample membership in multiple assays, using the upset algorithm in UpSetR
Source: R/upsetSamples.R
upsetSamples.RdCreate a generalized Venn Diagram analog for sample membership in multiple
assays, using the upset algorithm in UpSetR
Usage
upsetSamples(
MultiAssayExperiment,
nsets = NULL,
sets = names(MultiAssayExperiment),
nintersects = NA_integer_,
order.by = "freq",
check.names = FALSE,
...
)Arguments
- MultiAssayExperiment
A
MultiAssayExperimentobject- nsets
numeric(1)The number of sets to analyze. If specified,setswill be ignored.- sets
character()A character vector of names in MultiAssayExperiment to use. If specified,nsetswill be ignored.- nintersects
numeric(1)The number of intersections to plot. By default, all intersections will be plotted.- order.by
How the intersections in the matrix should be ordered by. Options include frequency (entered as "freq"), degree, or both in any order.
- check.names
logical(1)Whether to munge names as in thedata.frame()constructor (default FALSE).- ...
parameters passed to UpSetR::upset
Note
This function is intended to provide convenient visualization of assay
availability configurations in MultiAssayExperiment instances. The
UpSetR::upset function requires data.frame input and has
many parameters to tune appearance of the result. Assay name handling is
important for interpretability.
Examples
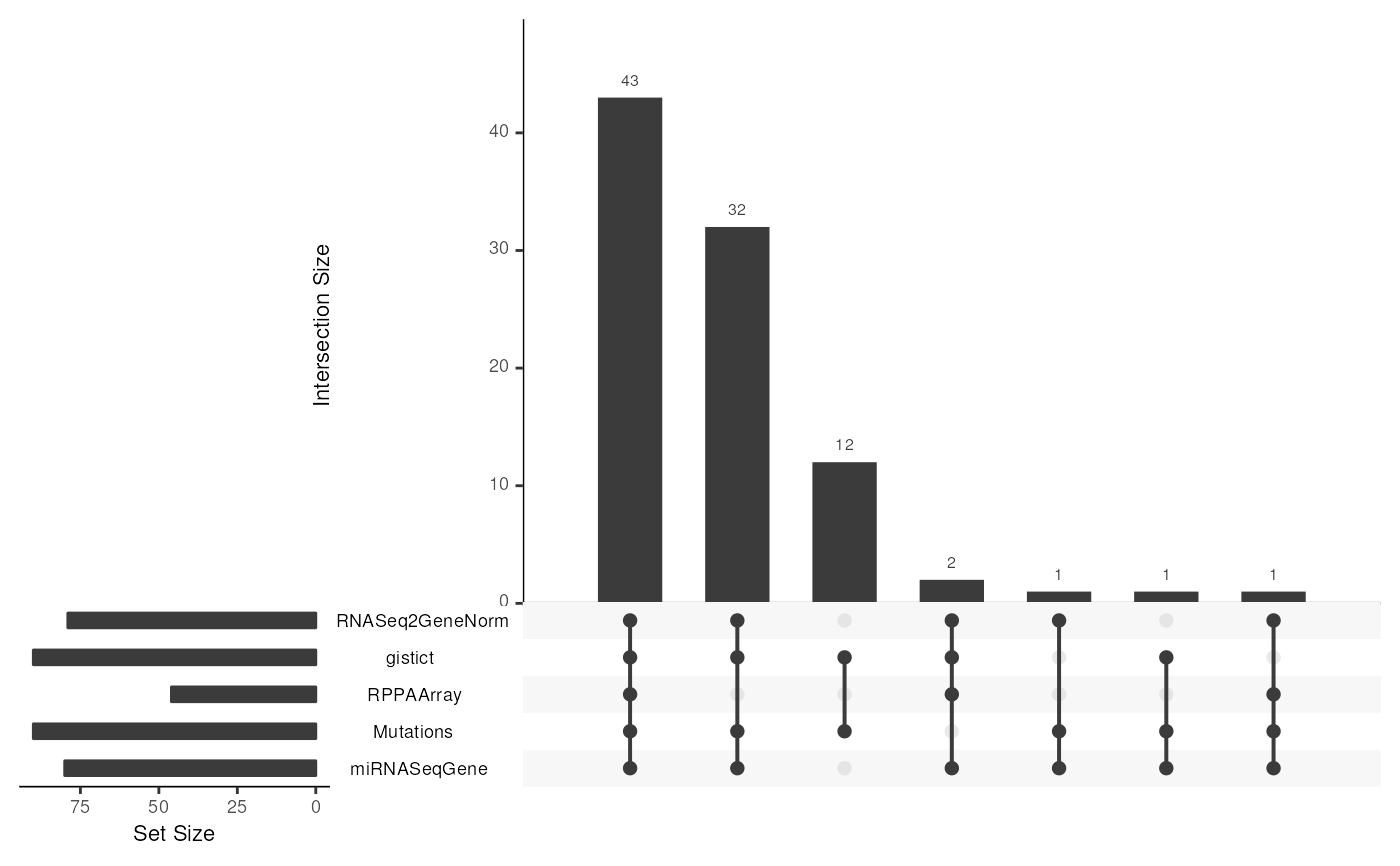
data(miniACC)
upsetSamples(miniACC)
#> Warning: `aes_string()` was deprecated in ggplot2 3.0.0.
#> ℹ Please use tidy evaluation idioms with `aes()`.
#> ℹ See also `vignette("ggplot2-in-packages")` for more information.
#> ℹ The deprecated feature was likely used in the UpSetR package.
#> Please report the issue to the authors.
#> Warning: Using `size` aesthetic for lines was deprecated in ggplot2 3.4.0.
#> ℹ Please use `linewidth` instead.
#> ℹ The deprecated feature was likely used in the UpSetR package.
#> Please report the issue to the authors.
#> Warning: The `size` argument of `element_line()` is deprecated as of ggplot2 3.4.0.
#> ℹ Please use the `linewidth` argument instead.
#> ℹ The deprecated feature was likely used in the UpSetR package.
#> Please report the issue to the authors.
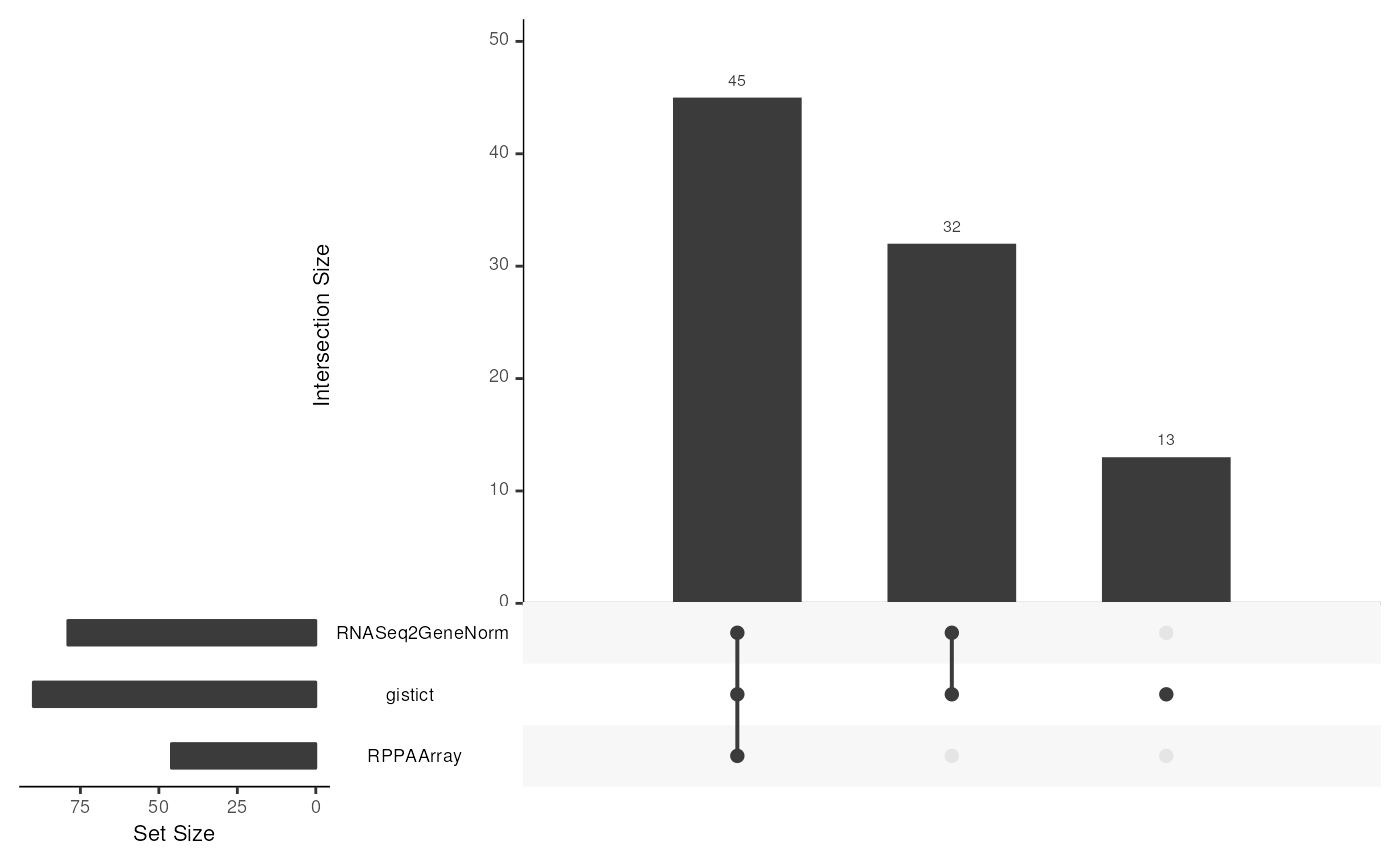
 upsetSamples(miniACC, nsets = 3, nintersects = 3)
upsetSamples(miniACC, nsets = 3, nintersects = 3)