Terra for paper
Introduction
This workspace provides a fully reproducible example of copy number variation (CNV) and single nucleotide variants (SNV) analysis of tumor samples without matching normal profile, described in the recent publication [link].
Reliable analysis of clinical tumor-only whole exome sequencing data
Oh et al., JCO Clin Cancer Inform. 2020 Apr;4:321-335. doi: 10.1200/CCI.19.00130.
Benefits
The major benefits of having Terra workspace with research paper are:
1) Data storage, pipeline (compute-intense), and downstream analysis are all available in one place
2) Improved reproducibility
3) Sharing code and providing additional information not included in the paper are available through the workspace
Features
1. Data
- Paper used TCGA controlled data (BAM files) → Synthetic dataset for public workspace
- Public reference files (stored in GCP, some are directly available through Terra)
- Researcher’s own data (BED, COSMIC VCFs stored in GCP) → some available with ‘Requester pays’
2. Workflows
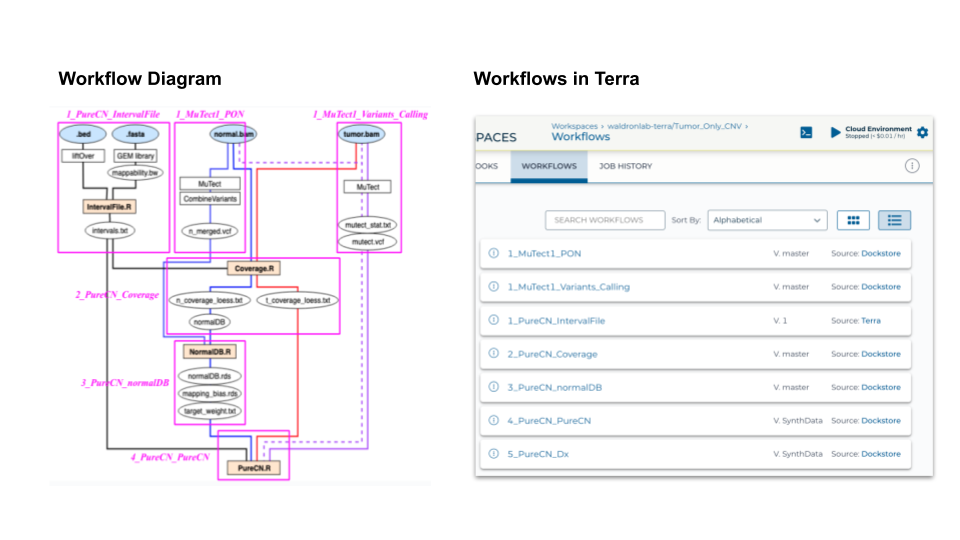
Left side of the diagram summarizes the CNV/SNV analysis workflow this paper used. Pink boxes around each segment of the pipeline are built into 7 WDL workflows in Terra, based on their modulatiry, input/output requirements. The numbered prefix represents the order each workflows should run. For example, 1_MuTect1_PON, 1_MuTect1_Variants_Calling, and 1_PureCN_IntervalFile can run all at the same time because they don’t have any dependence on other workflows for their inputs.
These workflows incorporate many different runtime environments (e.g. GATK, MuTect, Bioconductor,…).
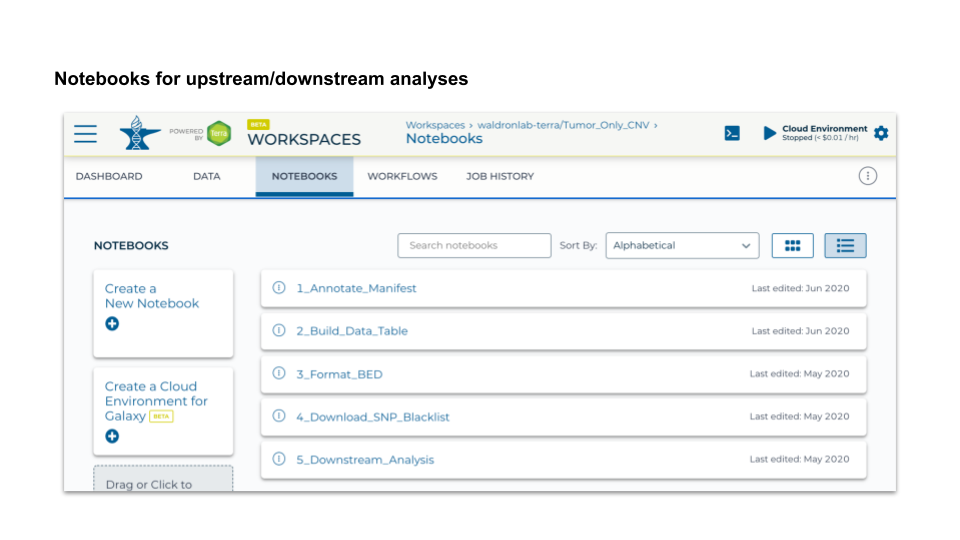
3. Notebooks
- 5 Jupyter notebooks using R → 4 for data pre-processing and 1 for downstream analysis
- AnVIL package enables a direct connection between ‘Data’ and ‘Notebooks’
Terra in the classroom
Find out how to use Terra/AnVIL for teaching R/Bioconductor HERE!
Control Billing
How to setup:
Need a Google Cloud Platform billing account
- Can use educational credits billing account
- Can use Onix billing account
- Can use regular billing account backed by credit card
Positives:
- Each student selects a compute environment with known cost per hour:
each student can select what they need
- Terra auto-suspend of notebook runtimes helped keep costs low
- Students only have to login, don’t have to set up billing
Negatives:
- Each student selects a compute environment with known cost per hour:
No way to identify an over-spending student or to limit what runtime they must use
- No cost breakdown per student
Notes:
- Billing is post-pay (you can find out how much has been spent with ~24h delay)